RATIONALE
Cancer is a disease of the genome, governed by principles of Darwinian evolution, whereby a cell acquires successive genetic or epigenetic alterations that lead to malignant transformation. The microenvironment is responsible for the selective pressures acting upon cells, for example immune predation, or resource availability (e.g. oxygen and nutrients provided through vascularization), and can in turn be remodeled by the cancer cell. Ecological interactions between a tumor and its environment are thus increasingly scrutinized to understand carcinogenesis and tumor progression.
AIMS
The cancer ecology and evolution project of the rare cancers genomics team is a transversal research program led by Dr. Nicolas Alcala that aims to build a theoretical and analytical framework of cancer formation and development. The project makes use of existing mathematical and computational models, as well as development of new methods, applying them to the multi-omic data generated within other team projects (lungNENomics, panNENomics, MESOMICS, and SARCOMICS). We focus on three topics :
- 1. Reconstructing the evolutionary history of tumors, and inferring the evolutionary forces at work, such as mutation and tumor demography (neutral and selection-driven growth). We use genomic and epigenomic data to infer the sequence of events that occurred in the tumor.
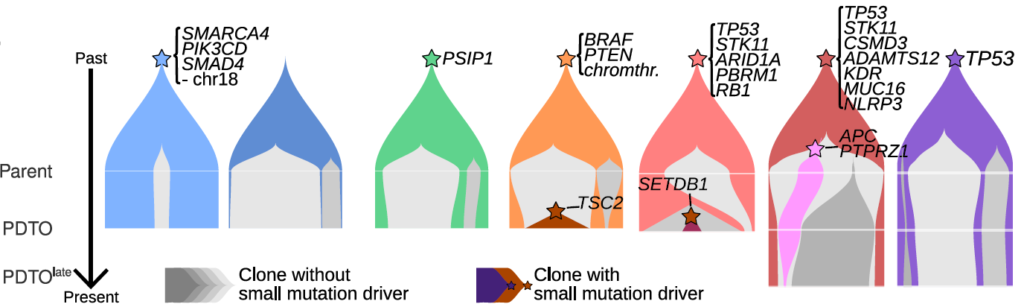
Temporal dynamics of tumor clones (colored shapes) in patient-derived tumor organoids (PDTO) of neuroendocrine neoplasms. Clonal complexity and selection (gray: neutrally evolving clones; colors: selected clones) increase with grade (left: low grade; right: high grade). From Dayton, Alcala et al. (Under review) https://www.biorxiv.org/content/10.1101/2022.10.31.514549v1
- 2. Understanding the interaction between tumor and microenvironment, and how they shape the selective landscape of tumors and their evolutionary trajectories. We couple transcriptomic, genomic, and histopathological data to infer tumor phenotypes, create a genotype-phenotype map and link it with the tumor microenvironment.
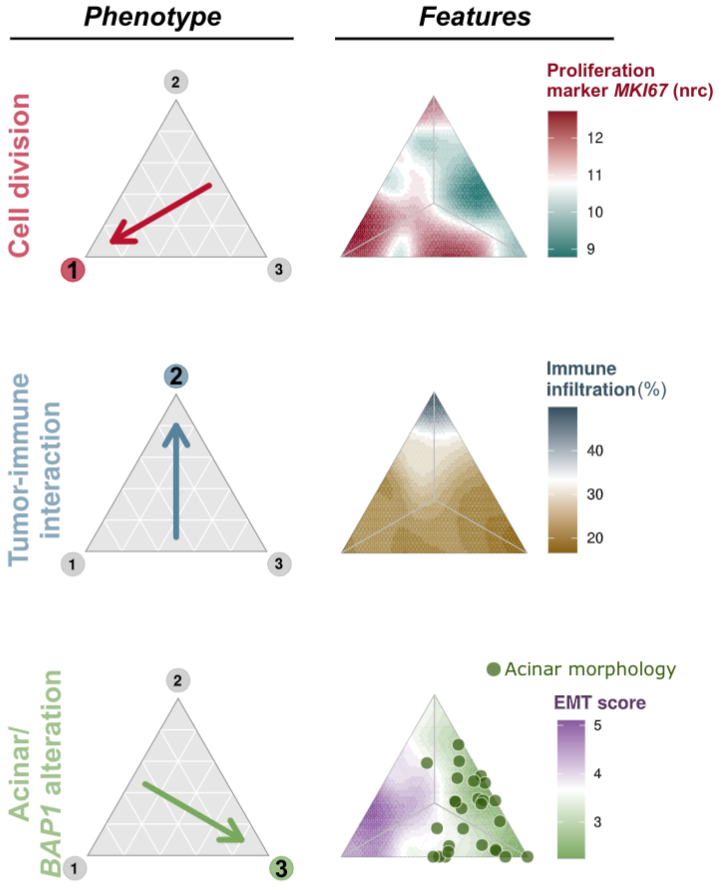
Malignant pleural mesothelioma phenotypes and associated features. Mesothelioma can be classified into three phenotypes (cell division, tumor-immune interaction, acinar), and each evolved under specific environmental conditions. From Mangiante, Alcala, Sexton-Oates, Di Genova et al., Nature Genetics (In press) https://www.biorxiv.org/content/10.1101/2021.09.27.461908v1
- 3. Developing statistical tools to quantify cell population evolvability, with the aim to anticipate malignant transformation, increased aggressiveness, and resistance to treatment. We use mathematical models to predict the behavior of certain quantities describing the intra-tumor heterogeneity of a population and mathematical statistics to design the best tool.
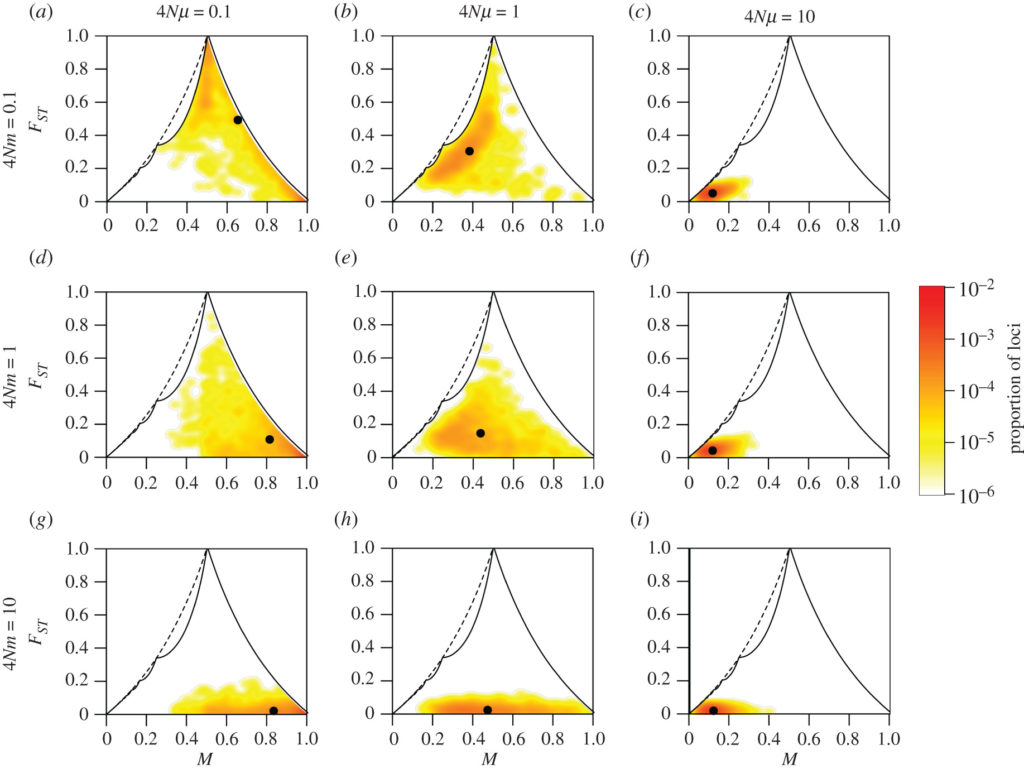
Computer simulations of a measure of differences in genetic composition (ranging from 0 for genetically similar populations to 1 for completely different populations), as a function of the frequency of genes M in two populations. Rows correspond to different rates of migration between the two populations (4Nm) and columns to different mutation rates for the individuals (4Nmu). From Alcala and Rosenberg, Philosophical Transactions of the Royal Society B https://royalsocietypublishing.org/doi/full/10.1098/rstb.2020.0414
FUNDING
Understanding the spatiotemporal eco-evolutionary interactions in malignant pleural mesothelioma. Worldwide Cancer Research (WCR, UK). Grant 24-0106. Coordinators. Active
Reconstructing the evolutionary history of neuroendocrine tumor subtypes. Neuroendocrine Tumor Research Foundation (NETRF, US). 2023 Mentored Award. Coordinators. Active
PhD fellowship to Laurane Mange. La Ligue contre le cancer. 2023-2026. Active
Population genetics of cancer evolution. France-Stanford Center for Interdisciplinary Studies (US). 2020-2021 Collaborative Research Project Grant. Active

